Agglomerative Clustering
This example covers the following:
- using the
BaseAlgoclass - validating search syntax
- converting parameters
- using
df_utilutilities - adding a custom metric to the algorithm
In this example, you will add scikit-learn's AgglomerativeClustering algorithm to the Splunk Machine Learning Toolkit. See the scikit-learn documentation for details on the AgglomerativeClustering algorithm.
In addition to inheriting from the BaseAlgo class, this example uses the convert_params utility and the df_util module. Additionally, you use scikit-learn's silhouette_samples function to create silhouette scores for each cluster label. See the scikit-learn documentation for more details on the silhouette_samples function.
This example uses the ML-SPL API available in the Splunk Machine Learning Toolkit version 2.2.0 and later. Verify your Splunk Machine Learning Toolkit version before using this example.
Steps
Do the following:
- Register the algorithm in
algos.conf.
- Register the algorithm using the REST API:
$ curl -k -u admin:<admin pass> https://localhost:8089/servicesNS/nobody/Splunk_ML_Toolkit/configs/conf-algos -d name="AgglomerativeClustering"
- Register the algorithm manually:
Modify or create thealgos.conffile located in$SPLUNK_HOME/etc/apps/Splunk_ML_Toolkit/local/and add the following stanza to register your algorithm[AgglomerativeClustering]
When you register the algorithm with this method, you will need to restart Splunk Enterprise.
- Register the algorithm using the REST API:
- Create the python file in the
algosfolder. For this example, you create$SPLUNK_HOME/etc/apps/Splunk_ML_Toolkit/bin/algos/AgglomerativeClustering.py
Ensure any needed code is imported. Import theconvert_paramsutility anddf_utilmodule.import numpy as np from sklearn.metrics import silhouette_sample from sklearn.cluster import AgglomerativeClustering as AgClustering from base import BaseAlgo from util.param_util import convert_params from util import df_util
- Define the class.
Inherit from theBaseAlgoclass:class AgglomerativeClustering(BaseAlgo): """Use scikit-learn's AgglomerativeClustering algorithm to cluster data."""
- Define the
__init__method.
- Check for valid syntax
- Convert parameters
- The
convert_paramsutility tries to convert parameters into the declared type. - In this example, the user will pass
k=<some integer>to the estimator -- however, when it is passed in via the search query, it is treated as a string. - The
convert_paramsutility will try to convert thekparameter to an integer and error accordingly if it cannot. - The
aliaslets users define the number of clusters withkinstead ofn_clusters.
- The
- Attach the initialized estimator to self with the converted parameters.
def __init__(self, options): feature_variables = options.get('feature_variables', {}) target_variable = options.get('target_variable', {}) # Ensure fields are present if len(feature_variables) == 0: raise RuntimeError('You must supply one or more fields') # No from clause allowed if len(target_variable) > 0: raise RuntimeError('AgglomerativeClustering does not support the from clause') # Convert params & alias k to n_clusters params = options.get('params', {}) out_params = convert_params( params, ints=['k'], strs=['linkage', 'affinity'], aliases={'k': 'n_clusters'} ) # Check for valid linkage if 'linkage' in out_params: valid_linkage = ['ward', 'complete', 'average'] if out_params['linkage'] not in valid_linkage: raise RuntimeError('linkage must be one of: {}'.format(', '.join(valid_linkage))) # Check for valid affinity if 'affinity' in out_params: valid_affinity = ['l1', 'l2', 'cosine', 'manhattan', 'precomputed', 'euclidean'] if out_params['affinity'] not in valid_affinity: raise RuntimeError('affinity must be one of: {}'.format(', '.join(valid_affinity))) # Check for invalid affinity & linkage combination if 'linkage' in out_params and 'affinity' in out_params: if out_params['linkage'] == 'ward': if out_params['affinity'] != 'euclidean': raise RuntimeError('ward linkage (default) must use euclidean affinity (default)') # Initialize the estimator self.estimator = AgClustering(**out_params)The convert_params utility is small and simple. When it is passed parameters from the search, they're received as strings. If you would like to pass them to an algorithm or estimator, you need to convert them to the proper type (e.g. an int or a boolean). The function does exactly this.
So when
convert_paramsis called, it will convert the parameters from the search to the proper type if they are one of the following:- float
- int
- string
- boolean
- Define the fit method.
- To merge predictions with the original data, first make a copy.
- Use the
df_util'sprepare_featuresmethod. - After making the predictions, create an output dataframe. Use the nans mask returned from
prepare_featuresto know where to insert the rows if there were any nulls present. - Lastly, merge with the original dataframe and return.
def fit(self, df, options): """Do the clustering and merge labels with original data.""" # Make a copy of the input data X = df.copy() # Use the df_util prepare_features method to # - drop null columns & rows # - convert categorical columns into dummy indicator columns # X is our cleaned data, nans is a mask of the null value locations X, nans, columns = df_util.prepare_features(X, self.feature_variables) # Do the actual clustering y_hat = self.estimator.fit_predict(X.values) # attach silhouette coefficient score for each row silhouettes = silhouette_samples(X, y_hat) # Combine the two arrays, and transpose them. y_hat = np.vstack([y_hat, silhouettes]).T # Assign default output names default_name = 'cluster' # Get the value from the as-clause if present output_name = options.get('output_name', default_name) # There are two columns - one for the labels, for the silhouette scores output_names = [output_name, 'silhouette_score'] # Use the predictions and nans-mask to create a new dataframe output_df = df_util.create_output_dataframe(y_hat, nans, output_names) # Merge the dataframe with the original input data df = df_util.merge_predictions(df, output_df) return dfThe prepare features does a number of things, and is just one of the utility methods in df_util.py.
prepare_features(X, variables, final_columns=None, get_dummies=True)This method defines conventional steps to prepare features:
- drop unused columns - drop rows that have missing values - optionally (if get_dummies==True) - convert categorical fields into indicator dummy variables - optionally (if final_column is provided) - make the resulting dataframe match final_columns
Args:
X (dataframe): input dataframe variables (list): column names final_columns (list): finalized column names - default is None get_dummies (bool): indicate if categorical variable should be converted - default is True
Returns:
X (dataframe): prepared feature dataframe nans (np array): boolean array to indicate which rows have missing values in the original dataframe columns (list): sorted list of feature column names
Output shape: In this example, you add two columns rather than just one column to the output. You need to make sure that the output_names passed to the create_output_dataframe method has two names in it.
Finished example
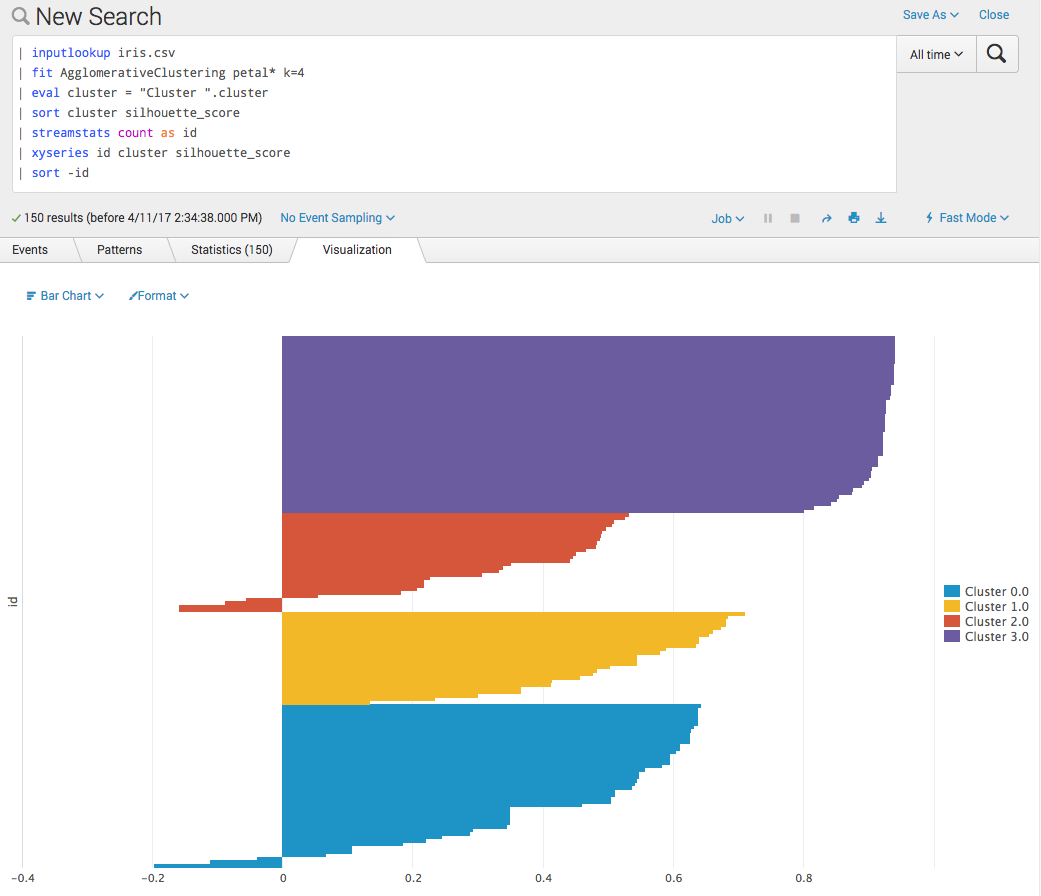
import numpy as np from sklearn.cluster import AgglomerativeClustering as AgClustering from sklearn.metrics import silhouette_samples from base import BaseAlgo from util.param_util import convert_params from util import df_util class AgglomerativeClustering(BaseAlgo): """Use scikit-learn's AgglomerativeClustering algorithm to cluster data.""" def __init__(self, options): feature_variables = options.get('feature_variables', {}) target_variable = options.get('target_variable', {}) # Ensure fields are present if len(feature_variables) == 0: raise RuntimeError('You must supply one or more fields') # No from clause allowed if len(target_variable) > 0: raise RuntimeError('AgglomerativeClustering does not support the from clause') # Convert params & alias k to n_clusters params = options.get('params', {}) out_params = convert_params( params, ints=['k'], strs=['linkage', 'affinity'], aliases={'k': 'n_clusters'} ) # Check for valid linkage if 'linkage' in out_params: valid_linkage = ['ward', 'complete', 'average'] if out_params['linkage'] not in valid_linkage: raise RuntimeError('linkage must be one of: {}'.format(', '.join(valid_linkage))) # Check for valid affinity if 'affinity' in out_params: valid_affinity = ['l1', 'l2', 'cosine', 'manhattan', 'precomputed', 'euclidean'] if out_params['affinity'] not in valid_affinity: raise RuntimeError('affinity must be one of: {}'.format(', '.join(valid_affinity))) # Check for invalid affinity & linkage combination if 'linkage' in out_params and 'affinity' in out_params: if out_params['linkage'] == 'ward': if out_params['affinity'] != 'euclidean': raise RuntimeError('ward linkage (default) must use euclidean affinity (default)') # Initialize the estimator self.estimator = AgClustering(**out_params) def fit(self, df, options): """Do the clustering & merge labels with original data.""" # Make a copy of the input data X = df.copy() # Use the df_util prepare_features method to # - drop null columns & rows # - convert categorical columns into dummy indicator columns # X is our cleaned data, nans is a mask of the null value locations X, nans, columns = df_util.prepare_features(X, self.feature_variables) # Do the actual clustering y_hat = self.estimator.fit_predict(X.values) # attach silhouette coefficient score for each row silhouettes = silhouette_samples(X, y_hat) # Combine the two arrays, and transpose them. y_hat = np.vstack([y_hat, silhouettes]).T # Assign default output names default_name = 'cluster' # Get the value from the as-clause if present output_name = options.get('output_name', default_name) # There are two columns - one for the labels, for the silhouette scores output_names = [output_name, 'silhouette_score'] # Use the predictions & nans-mask to create a new dataframe output_df = df_util.create_output_dataframe(y_hat, nans, output_names) # Merge the dataframe with the original input data df = df_util.merge_predictions(df, output_df) return dfSilhouette plot examples
You can now make a silhouette plot. These can be useful for selecting the number of clusters if not known a priori.
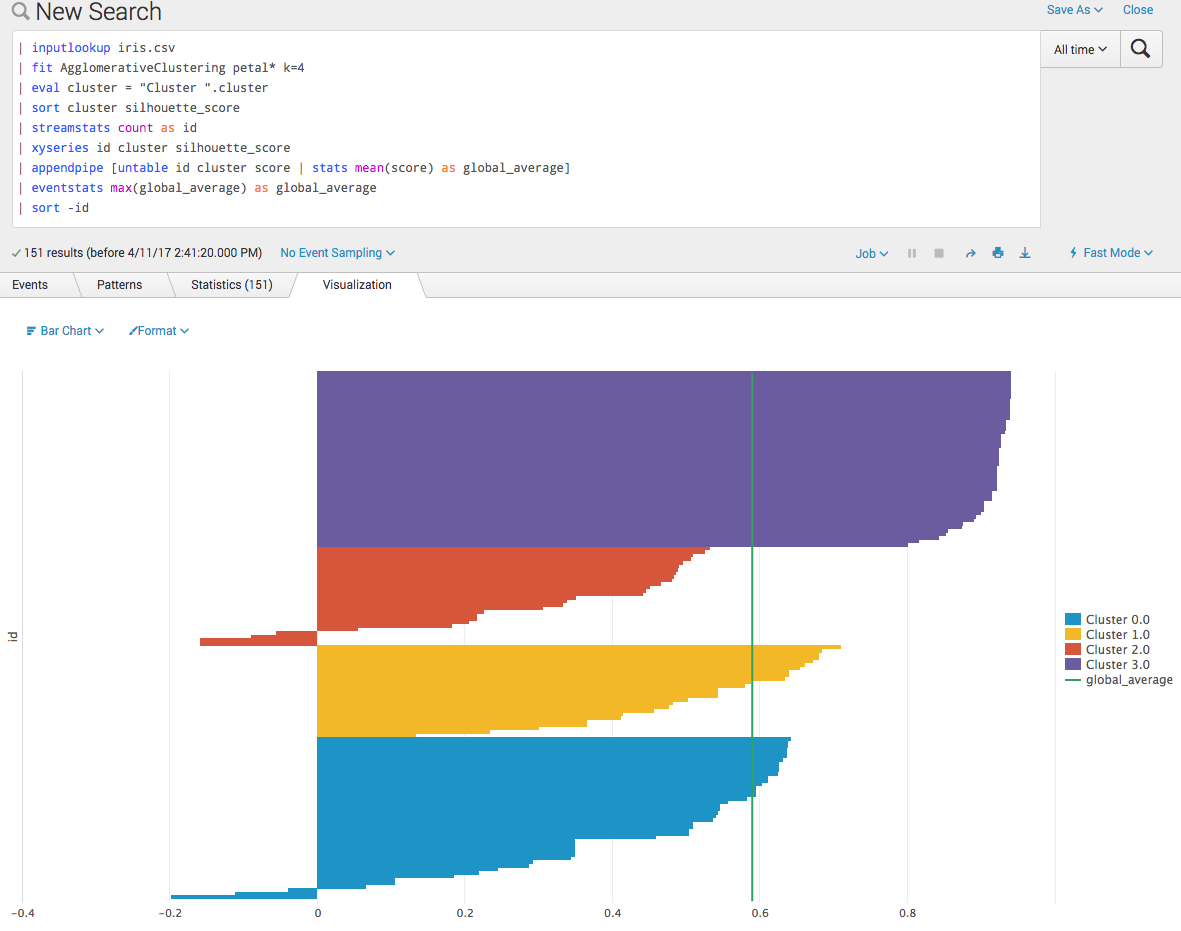
Often the global average is useful for such a plot. It is added in the following screenshot as a chart overlay:
| Correlation Matrix | Support Vector Regressor |
This documentation applies to the following versions of Splunk® Machine Learning Toolkit: 2.4.0, 3.0.0, 3.1.0, 3.2.0, 3.3.0, 3.4.0, 4.0.0, 4.1.0, 4.2.0, 4.3.0
 Download manual
Download manual
Feedback submitted, thanks!