Correlation Matrix example
You can add custom algorithms to the MLTK app. This example uses the Python library Pandas which is included with the Python for Scientific Computing (PSC) add-on.
PSC is a required add-on to use MLTK. PSC is a free add-on, available on Splunkbase.
The DataFrame.corr method constructs a correlation matrix. In addition to constructing the correlation matrix, you pass a parameter to the algorithm to switch between Pearson, Kendall and Spearman correlations. For more information on this method, see the Pandas library documentation: https://pandas.pydata.org/docs/
Add the Correlation Matrix algorithm
Follow these steps to add the Correlation Matrix algorithm. These steps fit a correlation matrix on all fields.
- Register the Correlation Matrix algorithm.
- Create the Python file.
- Define the class.
- Define the init method.
- Define the fit method.
Register the Correlation Matrix algorithm
Register the algorithm in algos.conf using one of the following methods.
Register the algorithm using the REST API
Use the following curl command to register using the REST API:
$ curl -k -u admin:<admin pass> https://localhost:8089/servicesNS/nobody/Splunk_ML_Toolkit/configs/conf-algos -d name="CorrelationMatrix"
Register the algorithm manually
Modify or create the algos.conf file located in $SPLUNK_HOME/etc/apps/Splunk_ML_Toolkit/local/ and add the following stanza to register your algorithm:
[CorrelationMatrix]
When you register the algorithm with this method, you must restart Splunk Enterprise.
Create the Python file
Create the Python file in the algos folder. For this example, you create $SPLUNK_HOME/etc/apps/Splunk_ML_Toolkit/bin/algos/CorrelationMatrix.py.
Import the relevant modules. In this case, use the BaseAlgo class which provides a skeleton class to catch errors:
from base import BaseAlgo
Define the class
Inherit from BaseAlgo. The class name is the name of the algorithm.
class CorrelationMatrix(BaseAlgo):
"""Compute and return a correlation matrix."""
Define the init method
The __init__ method passes the options from the search to the algorithm. Ensure that there are fields present and no from clause and that only valid methods are used by raising RuntimeError appropriately:
def __init__(self, options):
"""Check for valid correlation type, and save it to an attribute on self."""
feature_variables = options.get('feature_variables', {})
target_variable = options.get('target_variable', {})
if len(feature_variables) == 0:
raise RuntimeError('You must supply one or more fields')
if len(target_variable) > 0:
raise RuntimeError('CorrelationMatrix does not support the from clause')
valid_methods = ['spearman', 'kendall', 'pearson']
# Check to see if parameters exist
params = options.get('params', {})
# Check if method is in parameters in search
if 'method' in params:
if params['method'] not in valid_methods:
error_msg = 'Invalid value for method: must be one of {}'.format(
', '.join(valid_methods))
raise RuntimeError(error_msg)
# Assign method to self for later usage
self.method = params['method']
# Assign default method & ensure no other parameters are present
else:
# Default method for correlation
self.method = 'pearson'
# Check for bad parameters
if len(params) > 0:
raise RuntimeError('The only valid parameter is method.')
The options that are passed to this method are closely related to the SPL search query being used.
For a simple query such as:
| fit LinearRegression sepal_width from petal* fit_intercept=t
The options returned are:
{
'args': [u'sepal_width', u'petal*'],
'params': {u'fit_intercept': u't'},
'feature_variables': ['petal*'],
'target_variable': ['sepal_width']
'algo_name': u'LinearRegression',
}
This dictionary of options includes:
- args (list) - a list of the fields used - params (dict) - any parameters (key-value) pairs in the search - feature_variables (list) - fields to be used as features - target_variable (list) - the target field for prediction - algo_name (str) - the name of algorithm
Other keys that may exist depending on the search:
- model_name (str) - the name of the model being saved ('into' clause)
- output_name (str) - the name of the output ('as' clause)
The feature_fields and target field are related to the syntax of the search. If a from clause is present:
| fit LinearRegression target_variable from feature_variables
Whereas with an unsupervised algorithm such as KMeans:
| fit KMeans feature_variables
The feature_variables in the options have not been wildcard matched against the available data. If there are wildcards (*) in the field names, the wildcards are present in the feature_variables.
Define the fit method
The fit method is where you compute the correlations. And then returns the DataFrame.
def fit(self, df, options):
"""Compute the correlations and return a DataFrame."""
# df contains all the search results, including hidden fields
# but the requested requested are saved as self.feature_variables
requested_columns = df[self.feature_variables]
# Get correlations
correlations = requested_columns.corr(method=self.method)
# Reset index so that all the data are in columns
# (this is usually not necessary, but is for the corr method)
output_df = correlations.reset_index()
return output_df
When defining the fit method, you have the option to either return values or to do nothing, which returns None. If you return the dataframe, no apply method is needed. The apply method is only needed when a saved model must make predictions on unseen data.
End-to-end example
This Correlation Matrix example covers the following tasks:
- Using the
BaseAlgoclass - Validating search syntax
- Converting parameters
from base import BaseAlgo
class CorrelationMatrix(BaseAlgo):
"""Compute and return a correlation matrix."""
def __init__(self, options):
"""Check for valid correlation type, and save it to an attribute on self."""
feature_variables = options.get('feature_variables', {})
target_variable = options.get('target_variable', {})
if len(feature_variables) == 0:
raise RuntimeError('You must supply one or more fields')
if len(target_variable) > 0:
raise RuntimeError('CorrelationMatrix does not support the from clause')
valid_methods = ['spearman', 'kendall', 'pearson']
# Check to see if parameters exist
params = options.get('params', {})
# Check if method is in parameters in search
if 'method' in params:
if params['method'] not in valid_methods:
error_msg = 'Invalid value for method: must be one of {}'.format(
', '.join(valid_methods))
raise RuntimeError(error_msg)
# Assign method to self for later usage
self.method = params['method']
# Assign default method and ensure no other parameters are present
else:
# Default method for correlation
self.method = 'pearson'
# Check for bad parameters
if len(params) > 0:
raise RuntimeError('The only valid parameter is method.')
def fit(self, df, options):
"""Compute the correlations and return a DataFrame."""
# df contains all the search results, including hidden fields
# but the requested requested are saved as self.feature_variables
requested_columns = df[self.feature_variables]
# Get correlations
correlations = requested_columns.corr(method=self.method)
# Reset index so that all the data are in columns
# (this is necessary for the corr method)
output_df = correlations.reset_index()
return output_df
Example search
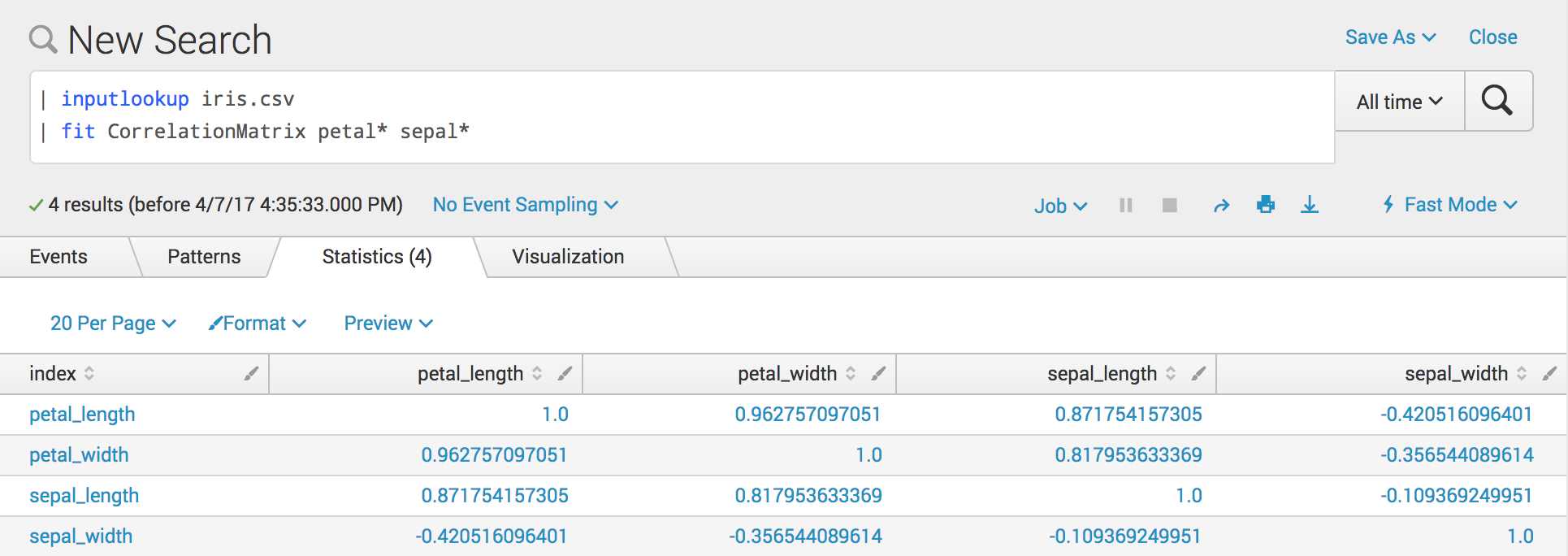
The following example search leverages the Correlation Matrix custom algorithm:
| inputlookup iris.csv
| fit CorrelationMatrix petal* sepal*
This image shows the example search in a Splunk search tab:
You might have to reorder your fields using the fields or table command.
| Package an algorithm for Splunkbase | Agglomerative Clustering example |
This documentation applies to the following versions of Splunk® Machine Learning Toolkit: 5.0.0, 5.1.0, 5.2.0, 5.2.1, 5.2.2, 5.3.0, 5.3.1, 5.3.3, 5.4.0, 5.4.1, 5.4.2, 5.5.0
 Download manual
Download manual
Feedback submitted, thanks!